 
ĭatabase search algorithm released in 2011 by Protein Metrics Inc. Developed by Jürgen Cox and others at the Max Planck Institute of Biochemistry. Identification of co-fragmented peptides improves the number of identified peptides. This combination enables analysis of large datasets on a desktop computer. Andromeda can function independently or integrated into MaxQuant.  
It can handle data with arbitrarily high fragment mass accuracy, it is able to assign and score complex patterns of post-translational modifications, such as highly phosphorylated peptides, and accommodates extremely large databases. On proteome data, Andromeda performs as well as Mascot, a widely used commercial search engine, as judged by sensitivity and specificity analysis based on target decoy searches. The former search takes place against a database containing all amino acid sequences assumed to be present in the analyzed sample, whereas the latter infers peptide sequences without knowledge of genomic data.Īndromeda is a peptide search engine based on probabilistic scoring. Peptide identification algorithms fall into two broad classes: database search and de novo search. In protein mass spectrometry, tandem mass spectrometry (also known as MS/MS or MS 2) experiments are used for protein/ peptide identification. In addition, hovering the mouse over the atom will highlight the applicable signal in the spectrum.ĭo not hesitate to contact us if you need any further piece of information.Further information: protein mass spectrometry Multiplet's boxes will be renamed with the corresponding atom number of the molecule and you will get an assignment label with the atom number over the applicable peak. 
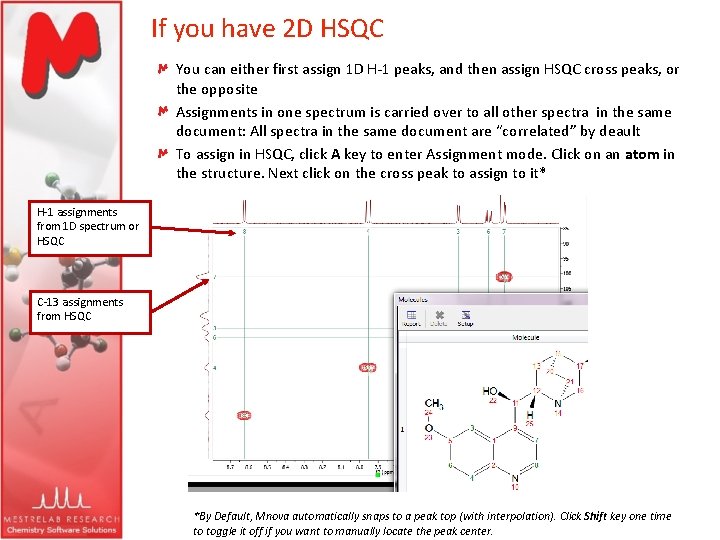
Mnova will automatically run a multiplet analysis and will fully assign your spectrum with the applicable molecular structure. You can also load your HSQC dataset in the same document, to increase the quality of the assignments. Just load your 1H-NMR spectrum with the applicable molecular structure and follow the menu 'Analysis/Auto Assignments'. Mnova provides a very simple interface to automatically assign your molecule. Please note that you need both Mnova NMR and Mnova NMRPredict Desktop in order to use Auto-Assignments. The Auto Assignment Algorithm of Mnova, combines several software techniques we had developed in recent years as tools for expert tasks such as automatic detection and characterization of spectral peaks, automatic solvent detection and automatic structure verification. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
